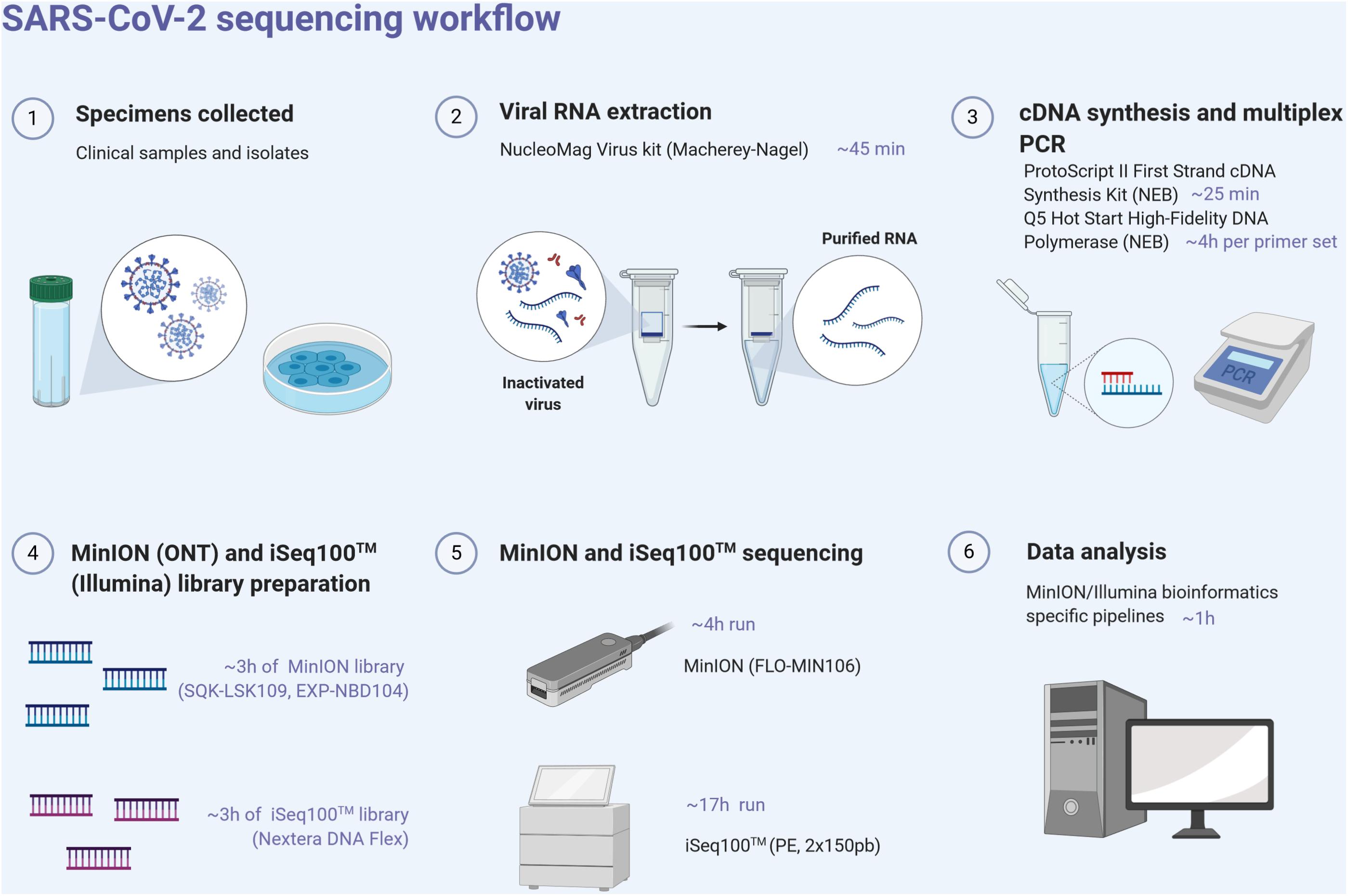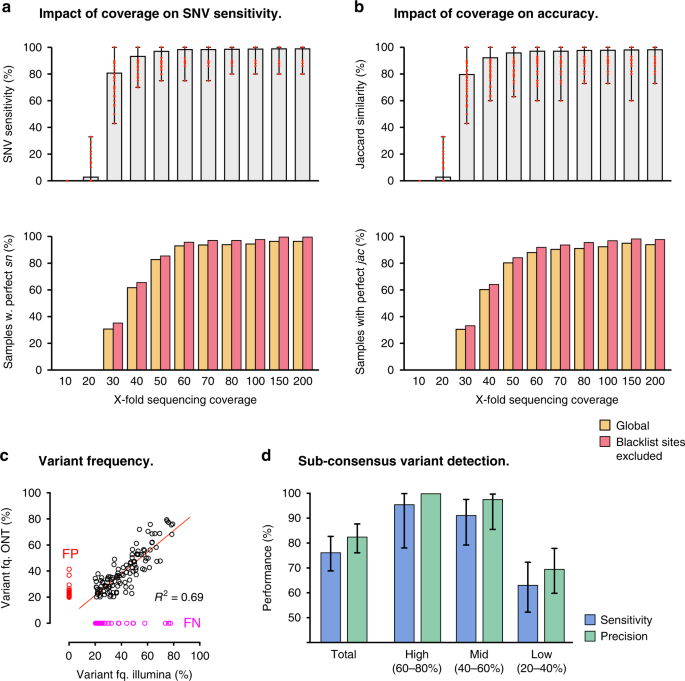
Analytical validity of nanopore sequencing for rapid SARS-CoV-2 genome analysis | Nature Communications

Evaluation of Nanopore sequencing for Mycobacterium tuberculosis drug susceptibility testing and outbreak investigation: a genomic analysis - The Lancet Microbe
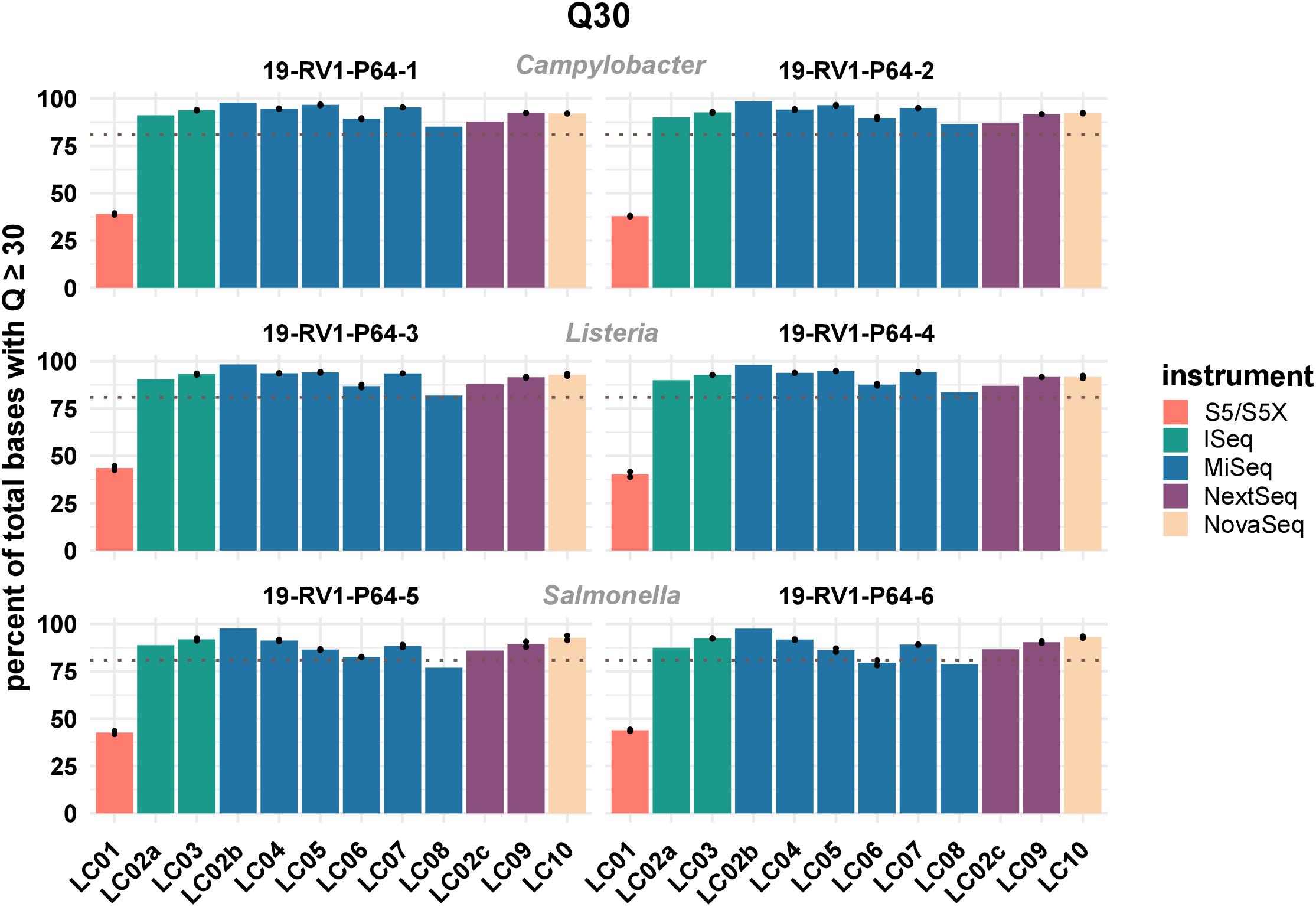
Frontiers | German-Wide Interlaboratory Study Compares Consistency, Accuracy and Reproducibility of Whole-Genome Short Read Sequencing
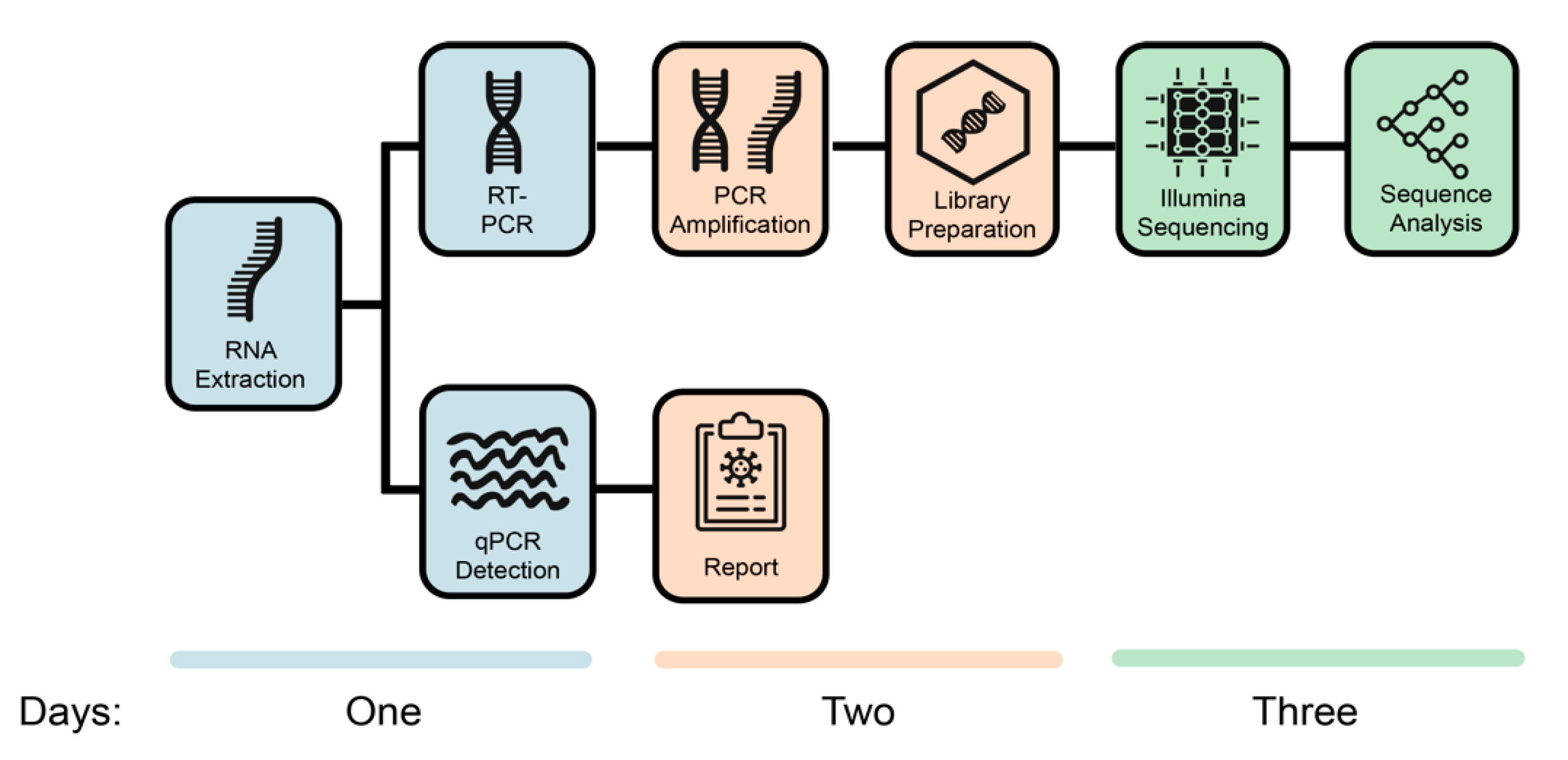
Genes | Free Full-Text | Whole Genome Sequencing of SARS-CoV-2: Adapting Illumina Protocols for Quick and Accurate Outbreak Investigation during a Pandemic
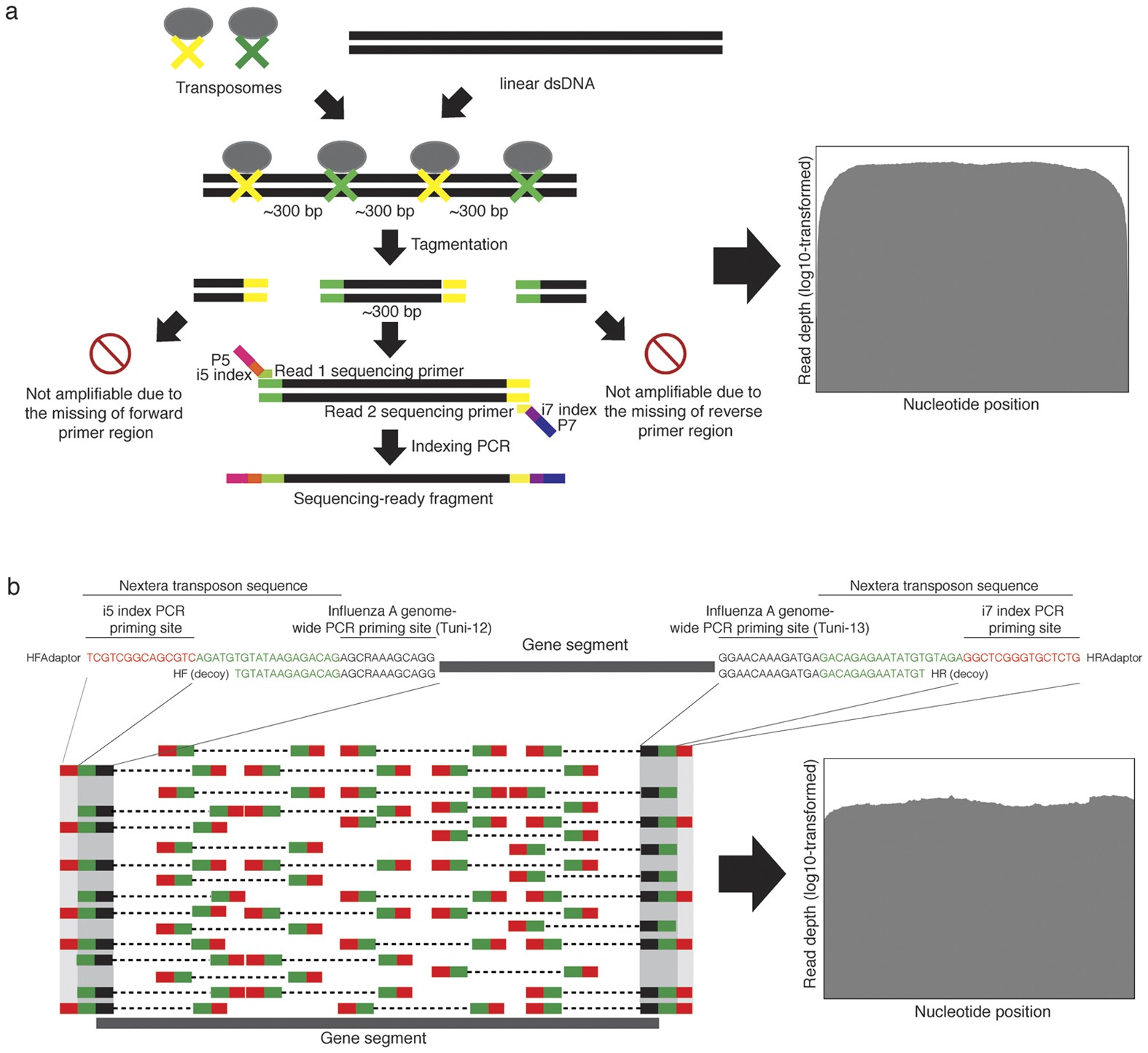
Contamination-controlled high-throughput whole genome sequencing for influenza A viruses using the MiSeq sequencer | Scientific Reports
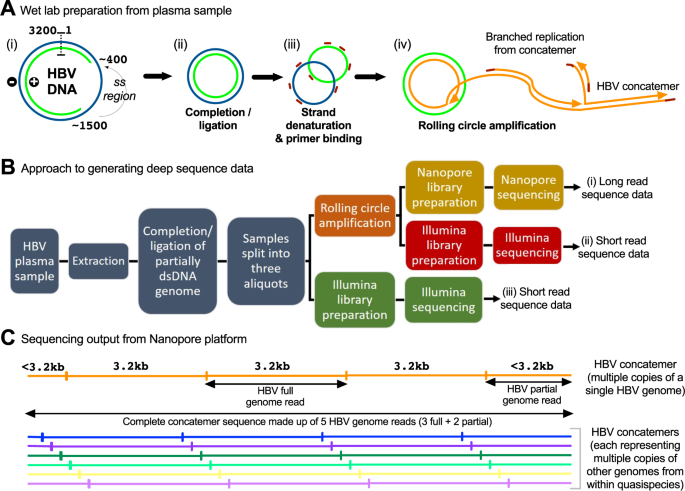
Illumina and Nanopore methods for whole genome sequencing of hepatitis B virus (HBV) | Scientific Reports
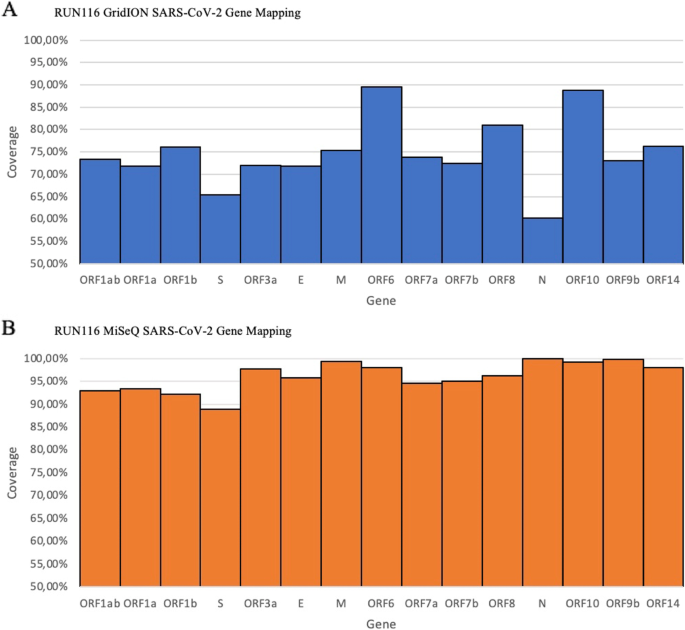
Comparison of SARS-CoV-2 sequencing using the ONT GridION and the Illumina MiSeq | BMC Genomics | Full Text